 
q INT Skip alignments with MAPQ smaller than INT samtools view -bq 1 file.bam > unique. Like for getting the unique reads (a single read mapping at one best position) I use: 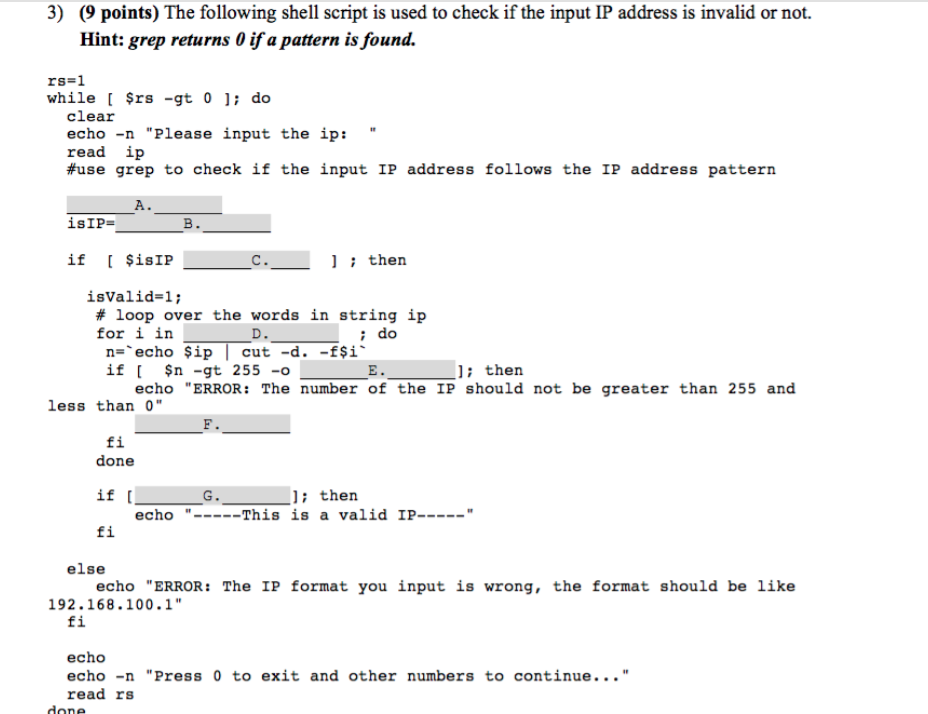
INT can be in hex in the format of /^0x+/ Įach bit in the FLAG field is defined as: Flag Chr DescriptionĠx0001 p the read is paired in sequencingĠx0002 P the read is mapped in a proper pairĠx0004 u the query sequence itself is unmappedĠx0010 r strand of the query (1 for reverse)Ġx0040 1 the read is the first read in a pairĠx0080 2 the read is the second read in a pairĠx0200 f the read fails platform/vendor quality checksĠx0400 d the read is either a PCR or an optical duplicate You can have grep read patterns from a file, one pattern per line, with the -f option: grep -x -F -A 1 -f File 2 File 1 Additionally,-F interprets patterns literally and not as regular expressions,-x only matches entire lines,-A N prints N lines following each match. bash grep Subject: read-messages cut -c10-80 Re: Linux suitable for. f INT Only output alignments with all bits in INT present in the FLAG field. Create a count file to extract the number of reads that maps within a gene. The sort INPUTFILE uniq -c sort -nr command string produces a frequency of. samtools view -b -F 4 file.bam > mapped.bamįrom the manual there are different int codes you can use with the parameter f, based on what you want: To get only the mapped reads use the parameter F, which works like -v of grep and skips the alignments for a specific flag. Heres an example of processing input as it is read. To get the output in bam, use: samtools view -b -f 4 file.bam > unmapped.bam It then reads the contents of the specified files (in the order specified), finds the lines that contain the search string, and finally returns those lines. Extracting Unique Elements from a List Problem You want to eliminate duplicate values from a list. To get the unmapped reads from a bam file use: samtools view -f 4 file.bam > unmapped.sam 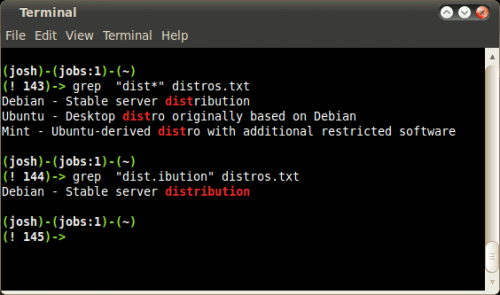
Hi, You get a bam (machine readable sam) file after mapping, and it contains information about mapped and unmapped reads. 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
